Formalin-fixed paraffin-embedded (FFPE) tissue samples are an important source of information for cancer researchers and are more readily available than frozen tumor samples collected from patients. These tissues allow for advancements in cancer treatment and prevention because they are backed with a wealth of clinical and outcome data enabling novel insights.
Now, researchers are able to more effectively analyze these FFPE tissue samples through the application of whole-genome sequencing (WGS) using next-generation sequencing (NGS) technology. NGS technology, however, has been restricted to frozen tumors. Using WGS for FFPE tissue samples is also made more difficult because of a number of factors from fragmented DNA and sequencing libraries with low complexity, to uneven genomic coverage and sequencing artifacts that are a result of the fixation method used.
To overcome these challenges, NGS of these samples require a high level of attempted sequencing coverage in order to compensate for the uneven coverage. WuXi NextCODE scientists Dr. Richard Williams and Dr. Shannon T. Bailey believe they have found the fix to this problem.
WuXi NextCODE SeqPlus
WuXi NextCODE’s SeqPlus is a nucleic acid extraction methodology that allows scientists and researchers to gather maximum value for FFPE sequencing compared to previous techniques. To understand more about SeqPlus, watch this free on-demand webinar from WuXi NextCODE , a genomic information company that uses sequencing data to improve people’s health around the globe.
FFPE tissues are most commonly sourced from archival material. During a patient’s course of treatment, FFPE samples are taken, stored, used and studied. It is inexpensive to create large samples and easy to maintain and preserve this library because they can be stored in a cabinet at room temperature.
However, proteins present in FFPE tissue samples are denatured during the fixing process, which can limit their application in certain types of immunohistochemistry studies. Furthermore, the nucleic acids in the FFPE tissue samples are not well preserved either and are often heavily fragmentated and limited in quantity.
WuXi NextCODE’s SeqPlus is capable of using original samples to unlock insights into previously-collected samples that have been of limited use due to restrictions of technology. They now have the technology to obtain high quality WGS read outs from FFPE samples. DNA extracted by the SeqPlus method results in at least 20 times the coverage for 98 percent of the genome. The advantages of this method include the ability to unlock new insights, supporting research into more targeted therapies, all while taking less time.
This method was validated using 15-year-old archival tissue samples that were provided by the National Cancer Institute (NCI). WuXi NextCODE’s SeqPlus was able to consistently confirm the presence of mutations in the DNA sequences from these samples which had been previously identified using traditional methods.
Xtalks spoke with Dr. Shannon T. Bailey, the Associate Director of Cancer Genetics at WuXi NextCODE, to understand the significance of these findings. “An important point that I want to make is that these are older samples. Most of what you’ll see with FFPE is that people typically look at younger samples — like a year or two [in age]. Going back 15 years is actually quite an accomplishment,” he said.
When discussing the importance of fixed tissues with Bailey, he said that the collection and fixation method of these tissues is particularly advantageous because scientists and researchers can use them almost indefinitely. In previous attempts to use WGS to analyze conventionally fixed tissues, the sequence needed to be looked at very deeply in order to make it work. But now more than ever, SeqPlus is becoming important in studying archival tissues.
“SeqPlus is a proprietary method for isolation of DNA as well as improvements in the library prep step of the sequencing workflow,” said Bailey. “So, what this method does is allow you to obtain high molecular weight DNA fragments.”
“Historically when you isolate DNA from FFPE tissues, the DNA is mostly fragmented into very small pieces, which we’ve learned doesn’t allow itself to be sequenced that well. But with SeqPlus, you extract much higher molecular weight fragments, and that is really the reason why we’re able to provide good sequencing coverage – the DNA is mostly intact,” said Bailey.
How It Was Validated
The goal of WuXi NextCODE’s study is to examine whether WGS of fixed esophageal squamous-cell carcinoma (ESCC) tissues stored for more than 10 years produces high quality sequencing results. The study included paired ESCC samples that had been stored for 10-16 years using a proprietary extraction and library preparation methodology, which uses WuXi NextCODE’s SeqPlus method.
The samples were collected from the Shanxi Province in China, where according to the National Mortality Survey, had the highest rates of esophageal cancer from 1973-1975. There were 20 FFPE samples and four alcohol-fixed samples in the collection.
In this study, however, there were no frozen tumor samples available to make a direct comparison of WGS results in both FFPE and frozen tissues. Nevertheless, various extended evaluations were made to analyze the outcome of the study. According to Dr. Alisa M. Goldstein, the principal investigator at NCI, they “assessed the SeqPlus performance of 20 tumor-germline paired older FFPE ESCC tissues, compared results to The Cancer Genome Atlas (TCGA) WGS frozen tumor study of esophageal cancer that was published a few years ago, and were also able to compare variant call concordance between WGS and a targeted, high-depth sequencing panel run at NCI/DCEG’s Cancer Genomics Research Lab (CGR).”
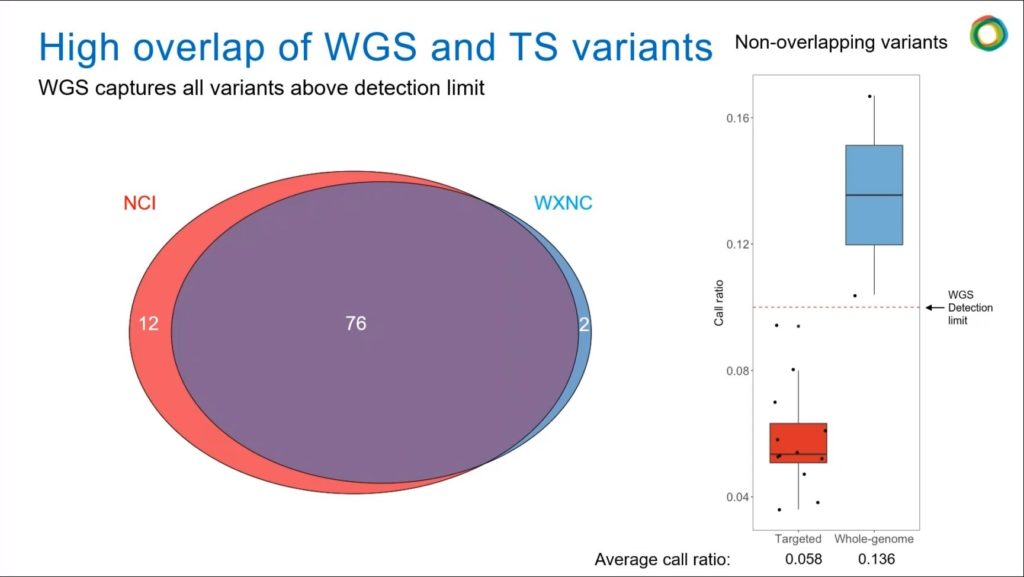
When looking at the samples, WuXi NextCODE wanted to make sure that they did not miss anything in terms of variants identified in the FFPE samples. Therefore, as seen above, they “compared the variants that were called by a high-depth targeted sequencing panel to those that were called using high-quality WGS,” said Bailey.
When this analysis was performed, the results showed that the WGS captured most of the variants that were called by the targeted panel, and only 12 were missed. This resulted in an 86 percent success rate in capturing the high-depth variants.
The 12 variants were then further studied, using the boxplots shown on the right of the figure above, showing the average call ratio to be less than 6 percent, whereas the detection limit of the WGS was 10 percent. This shows that they were able to capture everything that is possible using their WGS technology.
In the webinar, Bailey highlights some of the advantages of doing WGS in this method for this cohort. They were able to identify additional mutations in genes that were not covered in the panel, the variant cohorts match those that were previously published from other datasets, along with similar tumor mutation burden counts and copy number variant data.
New Insights from Archival Material
Goldstein underlines the findings of their study and said the aged fixed specimens had high molecular weight DNA for WGS and the SeqPlus methodology produced high-quality and consistent sequencing for all the FFPE tissue samples that were looked at. She said, “we believe that it unlocks the potential for massive-scale retrospective genomic analysis of stored pathology samples with associated clinical, therapeutic, and outcome data and provides opportunities to enable novel insights from these vast archival sample collections with concomitant, epidemiologic and clinical data.”
When using WuXi NextCODE SeqPlus with WGS, the method could help researchers develop more personalized therapies for patients. “We can get a more complete picture of a person’s own HLA components which is an important part of the immune make up of a person,” said Bailey.
In addition to identifying mutations, using SeqPlus to study cancer can also help identify important regions and drivers within the cancer genome, such as non-coding RNA, to provide a better understanding of the structural variations.
Dr. Shannon T. Bailey along with his collaborator Dr. Alisa M. Goldstein, share further insights into SeqPlus and WGS in this on-demand webinar.
This article was created in collaboration with the sponsoring company and the Xtalks editorial team.








Join or login to leave a comment
JOIN LOGIN