Biological Engineers at the Massachusetts Institute of Technology (MIT), have developed a simple method to customize bacteriophages – viruses that attack bacteria – to target specific pathogenic strains. Details of the process were published in the journal, Cell Systems.
While use of bacteriophages to fight bacterial infections has been proposed in the past, genetic engineering of the viruses to target a specific pathogen, has been traditionally expensive and time-consuming. The new system features a genetic template for a standard bacteriophage, with which scientists can add or subtract genes in order to customize the virus.
“These bacteriophages are designed in a way that’s relatively modular. You can take genes and swap them in and out and get a functional phage that has new properties,” said Timothy Lu, an associate professor of electrical engineering and computer science and biological engineering, and the principal investigator in the study.
Along with providing new treatment options for antibiotic resistant bacterial infections, the technology also has applications in the selection of beneficial bacteria in the gut. Of the trillions of bacteria that colonize the human digestive tract, many form symbiotic relationships which benefit the bacteria, as well as the host.
When conventional antibiotics are used to treat a pathogenic stomach infection, the good microbes are wiped out along with the bad, which can lead to further health problems. By using targeted bacteriophages to eliminate only the problem bacteria, the microflora balance of the gut is left undisturbed.
According to Lu, “Antibiotics can kill off a lot of the good flora in your gut. We aim to create effective and narrow-spectrum methods for targeting pathogens.” Lu explains that, “In the longer term you could design a specific phage that kills that bug but doesn’t kill the other ones, but more information about the microbiome is needed to effectively design such therapies.”
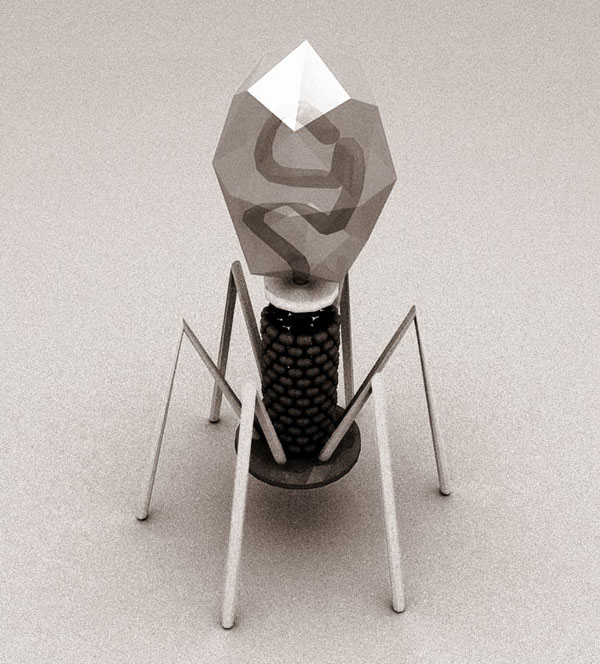
Lu and his team began to develop the technique using the T7 family of phages, whose natural target is Escherichia coli bacteria. By manipulating some of the genes encoding the tail of the bacteriophage – the region responsible for target specificity – they were able to change the bacterial strain targeted by the phage.
Lu said, “You keep the majority of the phage the same and all you’re changing is the tail region, which dictates what its target is.” Currently, there are no approved bacteriophages for human use. The only FDA-approved bacteria-targeting-viruses are used in the food production industry as an antimicrobial to prevent Listeria contamination.
Traditional methods of isolating naturally-occurring bacteriophage – including isolation from soil and sewage – is what makes the potential therapeutic method so time-consuming. Researchers also face another challenge in identifying and culturing the phages; different types of viruses have radically different life cycles and genome complexity. These issues have made phage therapy difficult to apply in clinical situations, in the past.
Lu and his colleagues hope that by providing a standardized genetic scaffold – along with the ability to ‘mix-and-match’ particular sequences – the technique could make bacteriophages a valid option for treating difficult bacterial infections.
The researchers used cultured yeast cells in order to do the viral genome manipulations. According to Lu, “Once we had that method, it allowed us very easily to identify the genes that code for the tails and engineer them or swap them in and out from other phages. You can use the same engineering strategy over and over, so that simplifies that workflow in the lab.”
To date, the team at MIT have successfully engineered the phages to target distinct strains of Gram-negative bacteria, including E. col, Klebsiella, and Yersinia. Gram-negative bacteria are particularly hard to treat with available antibiotics, and they can cause some of the most dangerous urinary, respiratory and gastrointestinal illnesses including pneumonia, sepsis, gastritis, and Legionnaires’ disease.
Sources:
- Customizing viruses to fight selected bacteria – http://www.medicalnewstoday.com/articles/299962.php
- Ando, H., Lemire, S., Pires, D., Lu, T. (2015). Engineering Modular Viral Scaffolds for Targeted Bacterial Population Editing. Cell Systems.










Join or login to leave a comment
JOIN LOGIN