For a while, it seemed like all microorganisms or pathogens were no match for antibiotics. But for the last several decades the appearance of deadly, antibiotic-resistant superbugs has caused doctors, researchers and government officials to reel back on prescribing these drugs. Today, around 700,000 people die from a drug resistance-related cause.
Despite measures to control antimicrobial resistance, scientists have yet to fully understand how these microorganisms become immune to the most potent antibiotic drugs.
Now, researchers from McMaster University in Ontario have revealed the secret to bacteria’s success. To do so, the team had to take a closer look at how antibiotics interact with bacteria.
“The antibiotics kind of poke into [the bacteria cell’s] membrane and stab the cell to death,” said Dr. Maikel Rheinstadter, lead author of the Nature Communications Biology publication and physics professor at McMaster University.
Dr. Rheinstadter and his team developed a technique to visualize this process, using polymyxin B, a potent antibiotic, and models of Gram-negative bacterial membranes.
In a two-step process, the antibiotics are electrostatically attracted to the bacterial cell membrane, adhering to the surface and penetrating into the cell itself. In the case of resistance, bacteria appear to manipulate their cell membranes to become less electrostatically attractive and more rigid, thwarting the efforts of the antibiotic drug.
First author and biochemistry undergraduate student, Andree Khondker said, “For the drug, it’s like going from cutting Jell-O to cutting through rock.”
Polymyxin B-resistant bacteria express a gene called mcr-1 which codes for an enzyme that modifies a fat molecule called lipid A in bacterial cell membranes. The researchers’ model suggests that bacteria expressing mcr-1 and carrying a high concentration of lipid A molecules results in a greater degree of antibiotic resistance.
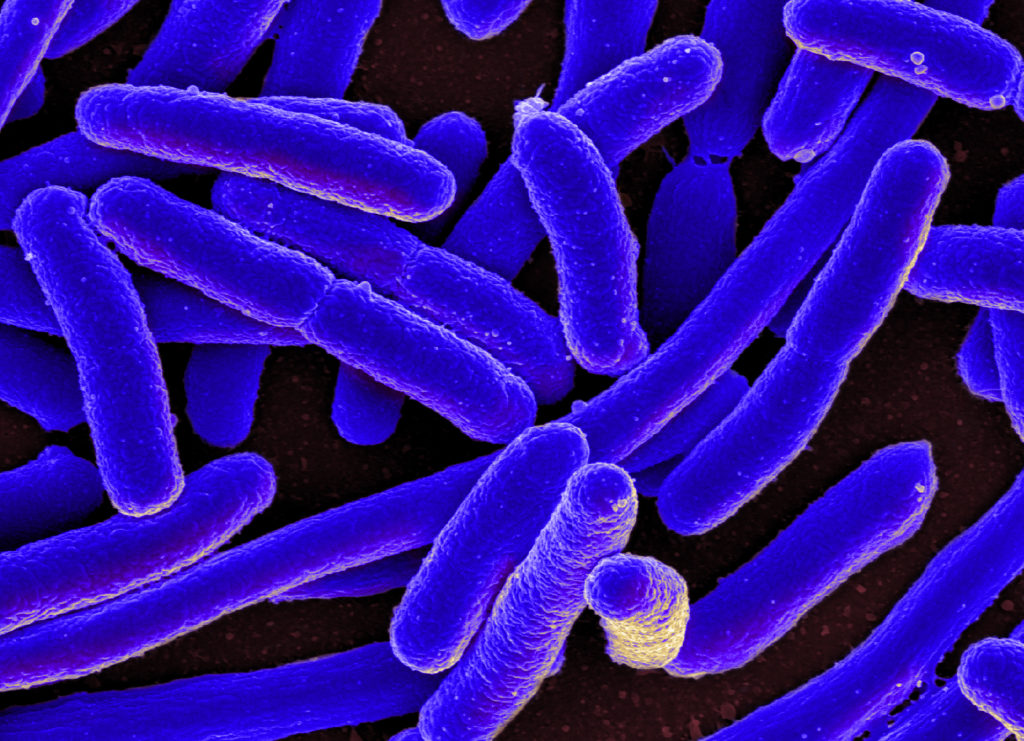
This helped to explain why antibiotic-resistant Escherichia coli – whose cell membrane contains 70 percent lipid A – showed an 11-fold greater relative membrane resistance.
In this study, the researchers evaluated resistance among Gram-negative bacteria such as E. coli and Pseudomonas aeruginosa. Other deadly strains like Staphylococcus aureus, considered one of the most notorious superbugs, may have alternative strategies to resisting antibiotic attack.
Still, this research can help scientists develop better therapies that can overcome bacterial defenses. In parallel, policymakers and governments are urging healthcare providers and pharmaceutical companies to combat antimicrobial resistance. For example, the UK government recently set goals to reduce antimicrobial use by 25 percent and reduce the number of healthcare-associated bloodstream bacterial infections by 50 percent, by 2024.
It won’t be enough for R&D to accelerate, but the way in which researchers tackle this problem will need to change. Dr. Rheinstadter’s findings were only made possible using X-ray diffraction, computer modeling and viewing the problem through the eyes of a physicist. It will take more research and innovation to fully understand and outsmart these resistant superbugs.




Join or login to leave a comment
JOIN LOGIN