Are you busy during this broadcast? See Broadcast #2 for an alternative time.
Advancement in cancer immunotherapy is associated with unraveling the complexities of immune suppressive mechanisms across different cancers. Quantification on multispectral multiplex-immunofluorescence (mIF) images allows detection of several biomarkers in a single section. In addition, new evidence using mIF techniques suggests that spatial analysis reveals novel insights in the tumor microenvironment. However, multispectral imaging is tile-based due to long scanning periods, which leads to insufficient data acquisition for significant spatial analysis. In this webinar, our featured speaker will present an automated workflow to study the spatial patterns of infiltrating cells in the tumor microenvironment based on multispectral mIF whole-slide scans. This was used to study the relationship between tumor proliferation and immune-response in non-small cell lung cancer (NSCLC) resections.
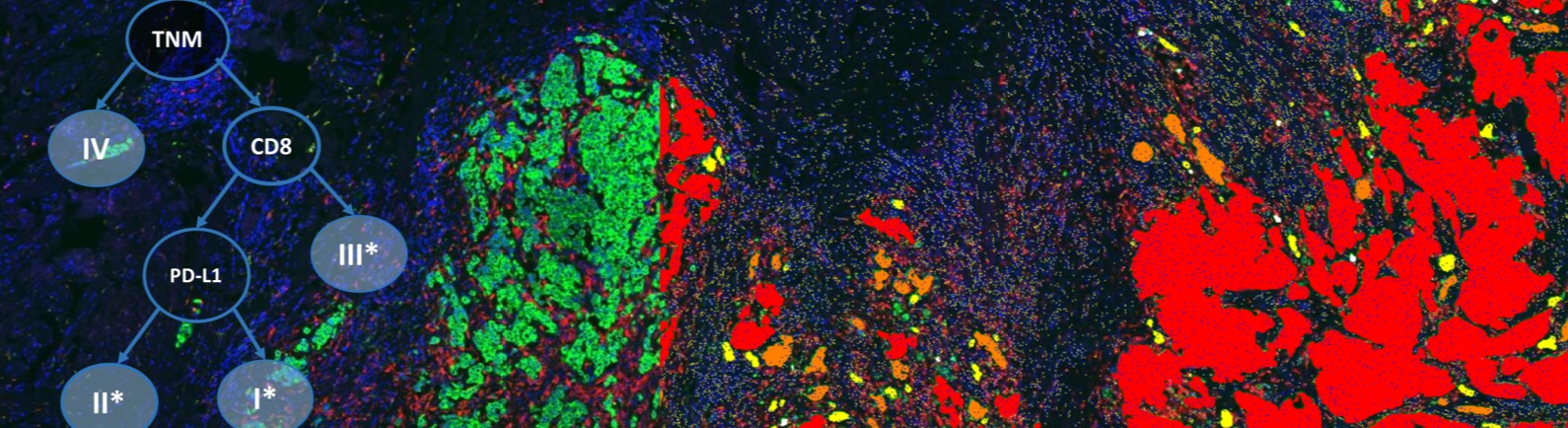
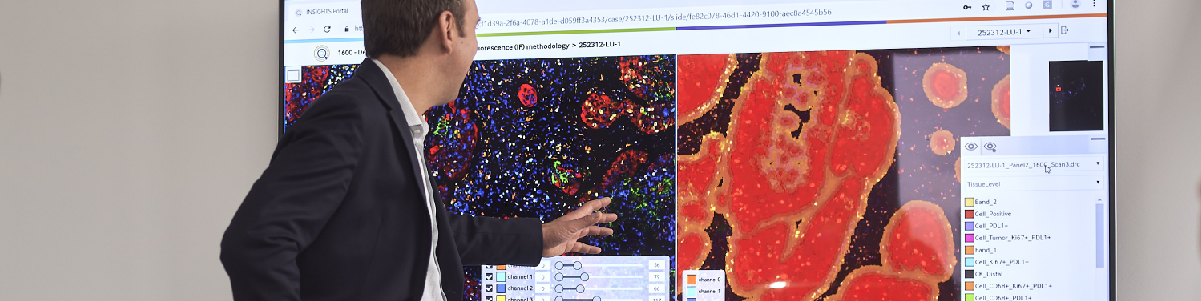
Formalin-fixed, paraffin-embedded NSCLC resection samples were stained by MedImmune with a custom-developed 7-plex mIF panel (CD68, CD8, Ki67, PD1, PD-L1, pancytokeratins-CK & DAPI) using the Opal method (PerkinElmer) in combination with an automated staining platform to ensure high throughput and consistent sample processing. Tiled scans were acquired with a Vectra Polaris (PerkinElmer) multispectral imaging system. Definiens developed a workflow which composes the tiled unmixed multispectral data to a whole-slide image and optimizes the layers for screen display and automated image analysis. Furthermore, images were shared on Definiens collaboration platform along with a chromogenic-IHC pseudocolor of the IF CK/DAPI signals and co-registered H&E section for pathologist annotations. These annotations were used in defining tumor center and invasive margin. The image analysis includes single-cell detection on the complete slide along with classification of subpopulations based on multi-marker positivity of individual cells. Part of the analysis is a high-quality tumor stroma separation based on the CK signal. The single-cell readouts were used to construct spatial biomarker expression patterns.
Speaker

Dr. Lorenz Rognoni, Senior Consultant Datafication, Definiens AG
Dr. Rognoni joined Definiens in 2015 and is currently leading the development of multiplex IHC image analysis (bright field and multispectral immunofluorescence) used in service projects. Before working in the field of digital pathology, he studied physics at the FU Berlin and LMU Munich and specialized in biophysics during his Ph.D. and postdoctoral research at TU Munich.
Who Should Attend?
This webinar will be ideal for pathologists and researchers in the biopharmaceutical industry. Other relevant areas include:
- Biomarker Development
- Drug Development
- Clinical Studies
- Imaging
- Histology
- Oncology
What You Will Learn
Learn about how Definiens developed an automated workflow for quantitative analysis of multispectral immunofluorescence whole-tissue scans. This allows the identification of spatial patterns for immunprofiling, where they could overcome the limitation of small regions of interest and provide a significant amount of data on the whole tumor region.
Xtalks Partner
Definiens
We improve patient lives by unlocking the tissue phenome.
In oncology, therapeutic strategies have shifted from a direct assault on cancer cells to recruiting the immune system for that purpose. Our mission is to accelerate breakthroughs for this approach by helping scientists leverage Tissue Phenomics to deepen understanding of disease biology and immune system mechanisms, to bring multi-omics data into a cancer-relevant context, and to facilitate the translation of new insights into novel therapies and treatment strategies. Our vision is to create unique patient profiles for an individualized standard of care, where patients experience fewer side effects and live longer.
Definiens’ Tissue Phenomics approach was awarded the 2013 Frost and Sullivan Company of the Year Award for Global Tissue Diagnostics and Pathology Imaging. For more information, please visit: www.definiens.com.
You Must Login To Register for this Free Webinar
Already have an account? LOGIN HERE. If you don’t have an account you need to create a free account.
Create Account