A team of researchers from Brown University have developed a new technique to map the three-dimensional forces that clusters of human cells exert on their surrounding environment.
This method could be helpful for scientists because it will allow them to understand how tissues form, how wounds heal and how tumors spread. In other words, this technique could be used for basic biology to clinical research or precision medicine.
“We know that the way groups of cells interact with their extracellular matrix is important, and we want to understand the instructions that tell these clusters to become organized into tissue-like architecture, or alternatively to become disorganized like an invasive tumor,” said Ian Y. Wong, an assistant professor at Brown University’s School of Engineering and corresponding author of a paper describing the work in the Proceedings of the National Academy of Sciences.
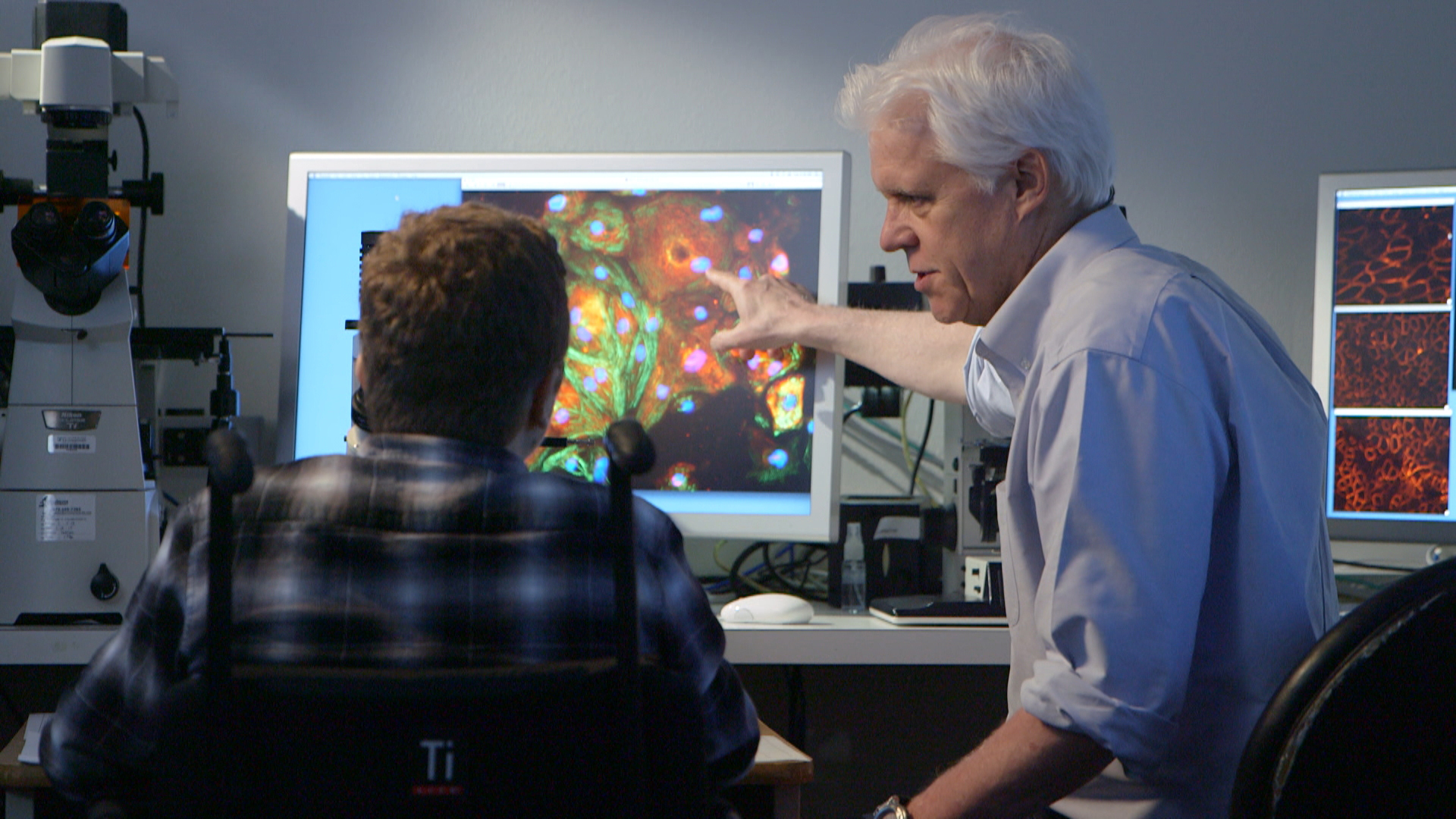
The technique uses traction force microscopy (TFM). This is an imaging method that is used widely to study the forces that are exerted by single cells. The way the TFM measurements work is that researchers place cells within biomaterials that mimic an extracellular matrix. This contains thousands of tiny fluorescent beads. Tracking the motion of beads as the cell moves inside the gel allows researchers to record the way in which the cell is pushing, pulling and twisting the biomaterial in 3-D.
“This technique gives us a way to profile these mechanical interactions between cells and matrix in a way that we couldn’t before,” said Dr. Wong.
The goal of this technique is to bring TFM to bear on multicellular clusters. Co-first author, Susan Leggett said, “we know that tumors, for example, tend to be spatially heterogeneous, with cells behaving differently throughout a tumor… so elucidating heterogenous behaviors across a multicellular cluster is something that’s important in a clinical context.”
To validate their method, researchers cultured clusters of mammary cells. They used a drug to stimulate epithelial to mesenchymal transition. This is a process where docile epithelial cells transform into elongated and highly mobile mesenchymal cells. Then, researchers found distinct force signatures for the different clusters (epithelial clusters, the mesenchymal clusters, and clusters that were in a transitory state). The team was then able to train a machine learning algorithm that could identify clusters from each group.
The team has made the code behind the technique freely available online in the hope that other researchers will find innovative ways to use it.








Join or login to leave a comment
JOIN LOGIN