A decade-long study that focuses on sequencing the genome of dozens of cancers has revealed how tumors form. This could change the way that scientists study cancer in hopes of finding better and more targeted treatment.
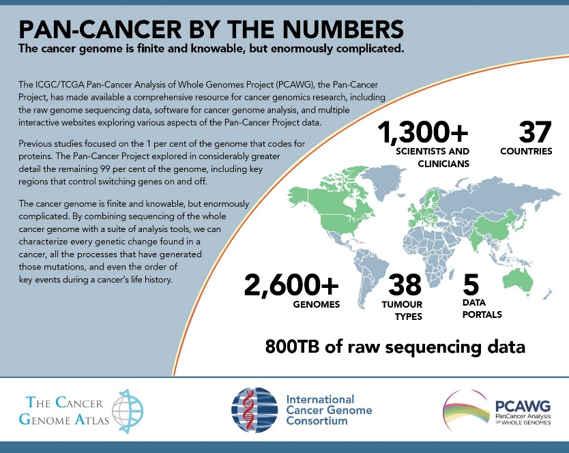
The Pan-Cancer Project brought together over 1,300 researchers to tackle this project. The study included 38 types of cancer and involved nearly 2,800 patients globally.

Nearly two dozen papers were published in Nature on Thursday, showing the results of the project, making it the largest and most comprehensive review of whole genome studies to ever exist.
Xtalks spoke with Dr. Lincoln Stein, head of adaptive oncology at the Ontario Institute for Cancer Research (OICR) and a member of the Pan-Cancer Project. He said, “over the last decade and a half, about 14 years, we’ve been able to sequence the genomes of individual cancers. Over that time, about 20,000 cancer genomes have been sequenced.”
Click play to listen in on the conversation
Previously, studies focus on only one percent of a genome that encodes proteins. This project, however, studies 99 percent of the genome. The Pan-Cancer Project has now made comprehensive resources available for cancer genomics research.
Dr. Stein said, “the problem with all the studies to-date has been that the sequencing is not the whole genome, it’s only the protein coding genes… while this has yielded a lot of information and in fact has really revolutionized how we diagnose and treat cancer, still, the protein coding portion of the genome is only one percent of the entire genome.”
“So, trying to paint a comprehensive portrait of the cancer with only one percent of the genome to work on, is like trying to put together a jigsaw puzzle that has tens of thousands of pieces and you only have one-hundredths of the pieces and you don’t have a box with a picture on it to guide you,” Dr. Stein said.
The Broad Institute calls this “the most comprehensive study of whole cancer genomes to-date, significantly improving our fundamental understanding of cancer and indicating new directions for developing diagnostics and treatments.”
Dr. Stein said that when looking at whole cancer genomes, a protein has a coding and non-coding portion, the latter of which is known among scientists as the dark matter, because they do not know much about it. They performed a uniform coordinated analysis that covered many different cancers, putting together over 2,600 cancers performing the whole genome sequencing. Then, the scientists took a few years to go through this dataset, creating the papers that were published in Nature last week.
Dr. Stein said, “it’s about 800 terabytes [of data] and I think that works out to a Netflix High Definition movie running continuously for 40 years.”
The data is shared through a cloud compute centre where researchers all around the world are able to access the files from their local cloud analysis center and run their analytical programs within that environment, giving them access to information instantly.
The way the project was organized, in terms of finding ways to make sense of the data, was by assigning scientists and placing them into a series of working groups that each look at different aspects of the cancer genome. For example, one group looked at cancer evolution and another looked at mutational patterns from environmental exposures. Once the results were compiled together and the series of simplified data was filed, a series of papers were produced to summarize the findings.
When speaking about the findings, Dr. Stein mentioned that one of the most interesting takeaways is that prior to the Pan-Cancer Project, when sequencing just the protein coding genes, scientists were able to find the causative mutation in two out of three patients. This means that 30 percent of the time there was nothing to find, and patients did not know why they had cancer. But now, they are able to read the mutation in 95 percent of patients, meaning that only 5 percent of patients do not have an explanation for their cancer. This resulted in a reduced failure rate from 30 percent to 5 percent.
Related: First of its Kind “Universal” Immune Cell Cancer Therapy
Other findings include being able to place mutations in chronological order, giving people a big opportunity to detect cancer at a much earlier stage and cure patients sooner. Furthermore, they did not find a large number of new cancer coding genes, which is good news because this means they are closing the catalog for cancer pathways. Finally, they found that each tumor type has a slightly different pattern of mutation in its genome and they can use that to create a machine learning system that uses AI to feed the whole genome of the tumor in order to tell what the tumor type is with accuracy.
The aim of the project, according to Dr. Stein, is to find the right therapy for each tumor. The challenge lies in the cancer itself. He explains that even if two people have the same tumor type under a microscope, are the same sex, and the same stage, one therapy might work with one person and not with the other. Stein says this is because there are fundamental parts of that cancer genome that we are not aware of.
“So we are trying to catalog each pattern of change in the cancer genome, and to understand how that particular pattern affects the tumors response to a particular therapy. In the not-so-distant future, the patient comes into the clinic, has his/her cancer sequenced and then the genome goes into a computer matching program, says, ‘well what other tumors have I seen that are similar to this and what was the clinical experience with patients who had this pattern?’”
In other words, they are trying to create a knowledge-based decision support system that allows clinicians to use genomic information from the patient’s tumor to select the therapy with the highest chances of success.
“This project is just one step along the way but it’s giving us the portrait of cancer, the lists of patterns, that allows us then to conduct the next series of studies in which we associate those patterns that we’ve observed in this study with response to therapy,” he said.





Join or login to leave a comment
JOIN LOGIN