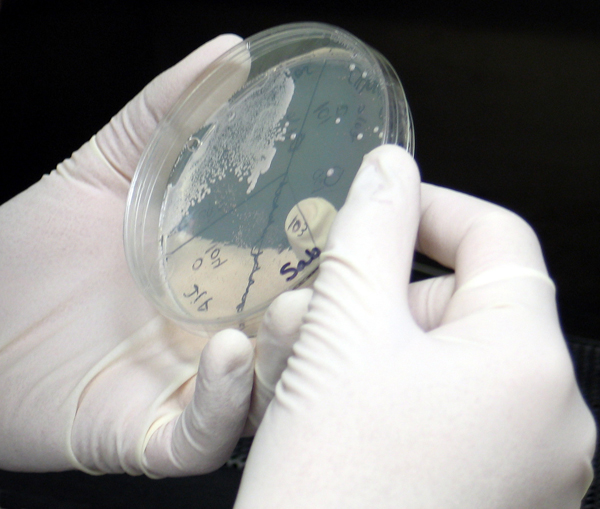
A global research team working on the Synthetic Yeast Project (Sc2.0), has assembled five new synthetic yeast chromosomes for the commonly-used Baker’s yeast (Saccharomyces cerevisiae). Working towards their goal of building synthetic versions of the microorganism’s 16 chromosomes, the team has achieved the goal of constructing a yeast strain with a 30 percent synthetic genome.
A set of seven papers on the subject, written by more than 200 authors, have been published in the journal, Science. The researchers sought to develop a better understanding of the yeast genome – which is often used in the lab as a model organism for human genetic research – by rebuilding it.
“This work sets the stage for completion of designer, synthetic genomes to address unmet needs in medicine and industry,” said Dr. Jef Boeke, director of NYU Langone’s Institute for Systems Genetics. “Beyond any one application, the papers confirm that newly created systems and software can answer basic questions about the nature of genetic machinery by reprogramming chromosomes in living cells.”
It’s been three years since the Sc2.0 team assembled the first synthetic yeast chromosome, which was composed of 272,871 base pairs. Five of the seven new publications describes each of the new yeast chromosomes, with one paper offering an overview of the research and the final paper detailing the 3D structures of the DNA in the yeast’s nucleus.
Building upon a wealth of previous work in yeast genetics, the researchers made strategic changes to the S. cerevisiae genome, including removing segments of DNA which are not believed to serve a functional role. Each of these individual changes gave rise to a genetically-altered yeast strain, which were organized into genetic libraries. Researchers then screened these libraries to identify variants with interesting or useful characteristics.
After identify which edits were most beneficial, the researchers combined the sequences into longer stretches of DNA. The team was able to improve this process by introducing the synthetic chunks of DNA into yeast cells, and manipulating the DNA replication machinery to complete the synthesis of the chromosome.
Previously, researchers had to build each chromosome bit-by-bit, making this process time-consuming and costly. The current study made multiple changes to this old protocol in an attempt to improve the parallel building of chromosomes.
The Sc2.0 project was truly a global effort, with researchers from different nations working together to build yeast strains with one or more synthetic chromosomes. “Steps can be accomplished at the same time in many locales and then assembled at the end, like networking laptops to create a global super computer,” said Dr. Leslie Mitchell, a post-doctoral fellow from Boeke’s lab at NYU Langone.
The technologies used by the Sc2.0 researchers will also be applied to Genome Project-Write (GP-write), a project aimed at synthesizing a complete human genome. Researchers contributing to the GP-write project hope to complete the goal in the next decade.










Join or login to leave a comment
JOIN LOGIN