Advances in molecular techniques such as next-generation sequencing have enabled scientists to resolve the complex genetic landscape of multiple cancers, down to specific mutations within tumor cells. These discoveries have led to new classifications systems, with each cancer subtype carrying its own diagnostic, prognostic and treatment implications.
Scientists have been able to harness this information to develop targeted cancer therapies, which are designed to act on cancer-specific genes, proteins or tissue environments. Much of the early work has been done with small molecule drugs, many of which are approved for the treatment of solid cancers such as breast cancer, colon cancer, lung cancer and melanoma. Thanks to targeted therapies, thousands of patients have access to lifesaving treatments that effectively stop their cancer from growing.
In principle, developing targeted therapies sounds straightforward. However, scientists face numerous challenges throughout all stages of drug development. For example, identification of the appropriate targets for drug development is dependent on the expression of the target in various tissues, the type of molecular test used and quality of data obtained. In the clinic, sponsors must consider best practices for recruiting patients and assembling a qualified team of investigators and clinicians to lead the study. Finally, when the drug is approved, the long-term impact of the new therapy on patient outcomes remains uncertain.
The complexities and challenges of developing targeted therapies are highlighted in the case of acute myeloid leukemia (AML), an aggressive blood cancer that is projected to kill an estimated 10,920 people in the US this year. Yet, over the last three years, the US Food and Drug Administration (FDA) approved eight targeted AML therapies — an unprecedented number reflecting both the urgency to find more effective treatments and the rapid pace of scientific advancement. Even so, the laboratory, operational and clinical challenges surrounding the development of targeted therapies remain formidable.
In a recent webinar, an expert panel from Medpace described the challenges and considerations for the clinical development of targeted therapies, using AML as a case study. The featured speakers described the merits and flaws of different next-generation sequencing tests and how they fit into the hematology-oncology lab, as well as the operational considerations for designing AML clinical trials. The speakers also discussed how the newly approved AML targeted therapies fit into the current treatment landscape.
Learn more about targeted therapies and AML by watching this on-demand webinar from Medpace, a global clinical contract research organization.
Using Next-Generation Sequencing to Identify Disease-Causing Mutations
In order to develop a targeted therapy, scientists require molecular knowledge of the cancer to match the therapy to the patient’s specific abnormality. To do this, scientists leverage next-generation sequencing (NGS), which encompasses a variety of tests that extract different types of genetic information from a DNA sample.
“When using NGS, there is always a trade-off,” says Dr. El Mustapha Bahassi, a research scientist at Medpace with over 20 years of clinical and research laboratory experience. “This trade-off will affect the cost, the time, the coverage and the read depth.”
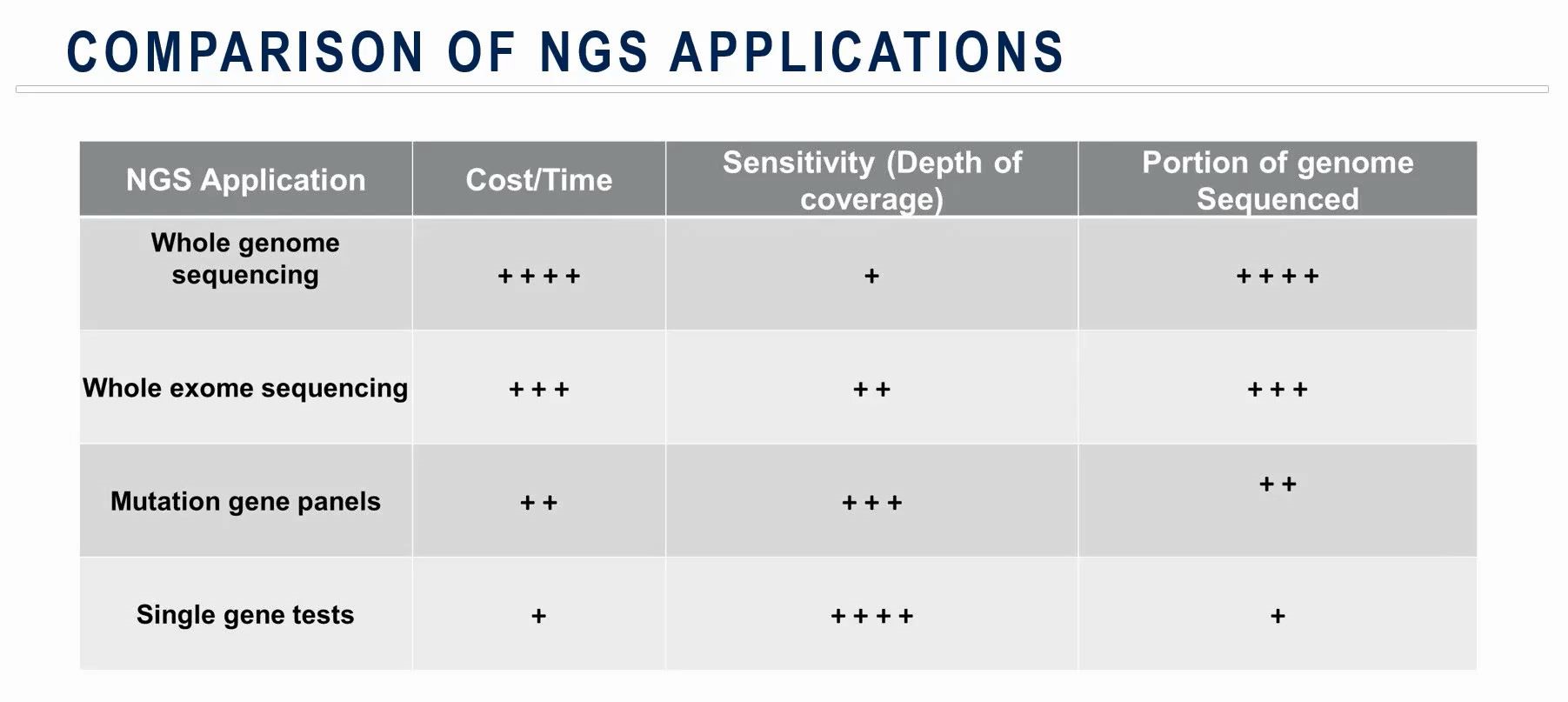
Whole-genome sequencing provides a holistic view of a person’s genome, but the coverage or depth of the reads is usually low. This means mutations that are present at very low allele frequency could be missed.
Whole-exome sequencing offers higher coverage compared to whole-genome sequencing at the expense of skipping non-protein-coding regions. While exons make up only one percent of the human genome, most disease-causing mutations affect these protein-coding genes, making this technique appealing to scientists.
Mutation gene panels and single gene tests only focus on a small portion of the genome but are superior in sensitivity (depth of coverage) and are often less costly.
Despite these tradeoffs, NGS is considered technically and clinically superior to non-NGS techniques. For example, NGS can identify actionable genomic alterations that could be missed by non-NGS techniques. This was the case when a team reported actionable genomic alterations in 65 percent of lung cancer tumors that were previously undetected using non-NGS methods. Moreover, many patients were then referred to clinical trials where a corresponding targeted therapy was being tested.
NGS has helped lay the framework for the molecular classification of AML, as demonstrated in a New England Journal of Medicine (NEJM) study published in 2013. Using DNA samples from 200 adult cases of de novo AML, the authors discovered a total of 23 significantly mutated genes plus another 237 that were mutated in two or more samples, as well as 118 gene fusions.
The authors categorized the mutated genes into functionally-related categories, which revealed many important biological relationships. This, in turn, provided a comprehensive foundation for an understanding of AML pathogenesis which then led to the discovery of many targeted therapies, Bahassi explains.
“Approximately 30 patients with AML have a FLT3 internal tandem duplication mutation, which is associated with the worst prognosis out of the known AML subtypes,” says Dr. Gregory Hale, senior medical director hematology and oncology at Medpace. According to the NEJM study, the FMS-related tyrosine kinase 3 (FLT3) gene, which codes for a cell-signalling protein found on hematopoietic cells, was mutated in over 50 samples.
The first tyrosine kinase inhibitor for AML, called midostaurin, was approved by the FDA in 2017. Gilteritinib, a more potent and more specific tyrosine kinase inhibitor was approved one year later, and at least two other FLT3 inhibitors are in clinical development. These agents are typically given in combination with other chemotherapy agents to boost their efficacy or to control disease as a post-remission maintenance therapy.
The frequency of these FLT3 mutations among AML patients may have contributed to the rapid development and approval of these targeted therapies. In fact, Hale says that midostaurin was “one of the first forays into targeted therapies for acute myeloid leukemia.”
Another gene implicated in AML is IDH, which codes for isocitrate dehydrogenase (IDH). This enzyme catalyzes the oxidative decarboxylation of isocitrate to alpha-ketoglutarate, a process involved in cellular metabolism. Mutations to the IDH gene results in a blockage of cell differentiation and are thought to contribute to cancer pathology.
“While FLT3 mutations are clearly associated with an inferior outcome, IDH mutations are considered prognostically neutral — they do not indicate an improved outcome or a worsened outcome, but they do represent a target for therapy,” says Hale.
Enasidenib, a drug that targets mutated IDH2 (an isoform of IDH), was recently approved based on a Phase I/II study that demonstrated an objective response rate of 40 percent.
Diagnosing AML
Data from these genetic analyses have helped inform the current guidelines for the diagnosis of AML. Two major institutions provide these guidelines: the European Leukemia Network (ELN) and the National Comprehensive Cancer Network. Both recommend genetic testing for patients with newly diagnosed AML, which includes conventional cytogenetics, screening for several gene mutations — including FLT3 — and testing for gene rearrangements.
“According to these two institutions, multiplex gene panels and NGS analysis may be used to obtain further information regarding prognosis, treatment decisions and eligibility for clinical trial participation,” explains Bahassi.
While clinicians may benefit from the swift turnaround time and relatively low cost of NGS, there are still challenges for implementation. For example, artifacts may arise during NGS sample preparation, such as mutations introduced during Polymerase Chain Reaction (PCR) amplification. For a hematology-oncology lab in particular, NGS is unable to discriminate between leukemia-related somatic mutations (acquired from disease) from pathogenic germline alterations. Finally, NGS has limited sensitivity for minimal residual disease assessment, which is an important step in risk stratification and treatment planning.
Many of these shortcomings can be addressed with deeper sequencing or additional testing by real-time PCR and flow cytometry. Still, there are certain mutations, such as the FLT3 internal tandem duplication that is not readily detected by NGS alone.
“There’s a need to use a parallel method to detect those duplications or develop new algorithms,” says Bahassi.
Nonmutation-Targeted Novel Agents
Somatic mutations are not the only drivers of disease. Sequencing studies have also revealed a host of abnormalities that can be treated with nonmutation-targeted therapies.
“Nonmutation-targeted novel agents are those that work through an epigenetic mechanism or that control or alter apoptosis,” explains Hale. “There are also agents that involve the targeted delivery of cytotoxic agents such as antibody-drug conjugates.”
Venetoclax is a potent inhibitor of BCL-2, an anti-apoptotic protein that has been found to play an oncogenic role in lymphoid and myeloid malignancies, including AML. Venetoclax is typically combined with hypomethylating agents such as azacytidine or decitabine to treat elderly AML patients. Recent studies demonstrated a composite complete response rate approaching 70 percent with a median overall survival approaching 18 months in elderly patients who were treated with this combination therapy.
“I would emphasize that when used alone as a single agent in the same population, the complete response rate is only six percent, with progression-free survival of 2.5 months,” says Hale. “So, there’s definitely a significant rationale for using venetoclax in combination with hypomethylating agents.”
Gemtuzumab ozogamicin is an antibody-drug conjugate comprising a humanized anti-CD33 and a derivative of the cytotoxic molecule, calicheamicin. The drug was initially approved in 2000 but was withdrawn following reports of relapse and severe adverse events. Subsequent pooled analysis of five randomized controlled trials demonstrated improvements in relapse rate and overall survival rate, leading to its reapproval in 2017.
It is unlikely that the FDA would approve a drug without a robust safety profile, but the example of gemtuzumab ozogamicin points to significant challenge in AML drug development. NGS tests have revealed dozens of new actionable targets, but there could be limited published data supporting the safety and efficacy of each. Antibody-drug conjugates, for example, are a relatively new class of biopharmaceutical drugs compared to traditional chemotherapeutic agents. The lack of published data may deter patients from participating in a clinical trial evaluating a new class of drugs.
Current State of AML Clinical Trials
Appealing to a patient’s interest is one of the many factors to consider when conducting an AML clinical trial. In addition to the amount of published data available, a patient’s interest is also influenced by the complexity of the protocol (i.e. complex or time-consuming procedures are undesirable) and whether or not there are support services available (e.g. educational materials, reimbursement for travel).
“Additionally, there are currently 541 actively recruiting trials for the treatment of patients with AML,” says Dr. Jeffrey Vassallo, associate director of clinical trial management at Medpace. “Both investigators and patients have many treatment options and this is further limited by the rapidly increasing smaller group of patients defined by molecular abnormalities.”
In other words, while the number of patients diagnosed with AML is big, the eligible patient population for a particular targeted therapy trial is small. To optimize trial success, Vassallo recommends a well-written protocol, supported by published data and the inclusion of procedures that would ideally support the study’s objectives without being too complex or burdensome to the patients involved. Additionally, sponsors should aim to select the right mix of experienced trial sites that support successful execution of the program within the intended timeframe.
Finally, the successful trial execution is predicated on relationships and expertise.
“It’s important to ensure that experts in the treatment of patients with AML are included at the investigative sites in addition to the sponsor and CRO, with an emphasis on the medical monitor, nurse practitioner, clinical trial manager, clinical research associates and an enthusiastic investigator,” notes Vassallo.
The approval of eight new targeted therapies in less than five years speaks to the progress in AML research.
“Certainly, the field is a moving target as we can see that ‘7+3’ chemotherapy regimen has been the induction regimen for 40 years,” says Hale. “Now, there’s venetoclax with hypomethylating agents that have demonstrated a markedly improved outcome and profile.”
He adds, “It is important to note that while targeted therapies demonstrate promising results in this patient population, none of these agents has yet achieved the clinical results that we have observed in acute myeloid leukemia M3 or in chronic myeloid leukemia with tyrosine kinase inhibitors.”
Regardless of the poor outcomes seen in patients with AML, disease recurrence continues to remain the primary cause of treatment failure for this patient population.
For further insights into the development of targeted therapies and advances in AML drug development, watch this on-demand webinar from Medpace.
This article was created in collaboration with the sponsoring company and the Xtalks editorial team.








Join or login to leave a comment
JOIN LOGIN