The Wyss Institute at Harvard University has revealed a new, at-home, rapid CRISPR COVID-19 test that can detect SARS-CoV-2 variants. The point-of-care test delivers results in about an hour and allows users to test for the virus using a simple saliva sample right at home.
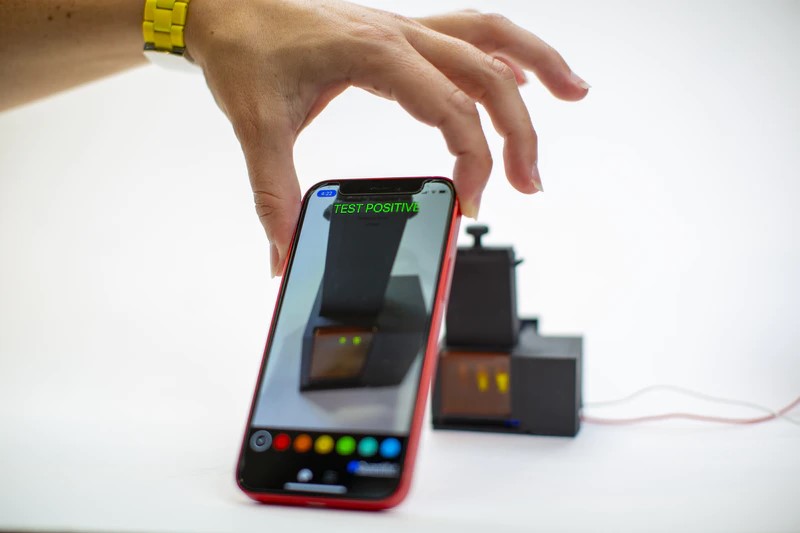
Results from the test are read and verified through a linked smartphone app within one hour.
The point-of-care COVID-19 diagnostic is called the minimally instrumented SHERLOCK (miSHERLOCK) and is based on CRISPR technology for the molecular detection of up to three different variants of SARS-CoV-2.
The test was developed by researchers at the Wyss Institute for Biologically Inspired Engineering at Harvard University, the Massachusetts Institute of Technology (MIT) and several Boston area hospitals. The researchers published the details of the device in a paper in Science Advances.
Related: LabCorp’s Innovative High-Speed COVID-19 Test Gets Emergency Use Authorization from the FDA
The researchers were able to successfully distinguish between three different variants of SARS-CoV-2 with the CRISPR COVID-19 test in experiments, and the device can be easily reconfigured to detect other variants, like Delta.
The research team opted to use saliva rather than nasopharyngeal swab samples as their diagnostic source material, as collecting saliva is far easier and faster for users at home. Moreover, studies show that saliva samples can contain a marginally higher SARS-CoV-2 viral load compared with nasopharyngeal samples (shown in a study involving hospitalized patients), and that the virus can be detected in saliva for a greater number of days post-infection.
However, unprocessed saliva contains enzymes that degrade various molecules, which can lead to a high rate of false positives. To overcome this challenge, the researchers developed a novel technique that involves addition of lysis buffer containing DTT and EGTA to the saliva and heating the sample to 95°C for three minutes to deactivate false positive-producing enzymes and also to open up any viral particles. A porous membrane was engineered to capture RNA on its surface.
“miSHERLOCK eliminates the need to transport patient samples to a centralized testing location and greatly simplifies the sample preparation steps, giving patients and doctors a faster, more accurate picture of individual and community health, which is critical during an evolving pandemic,” said co-first author on the paper Dr. Helena de Puig, a postdoctoral fellow at the Wyss Institute and MIT.
miSHERLOCK Workflow
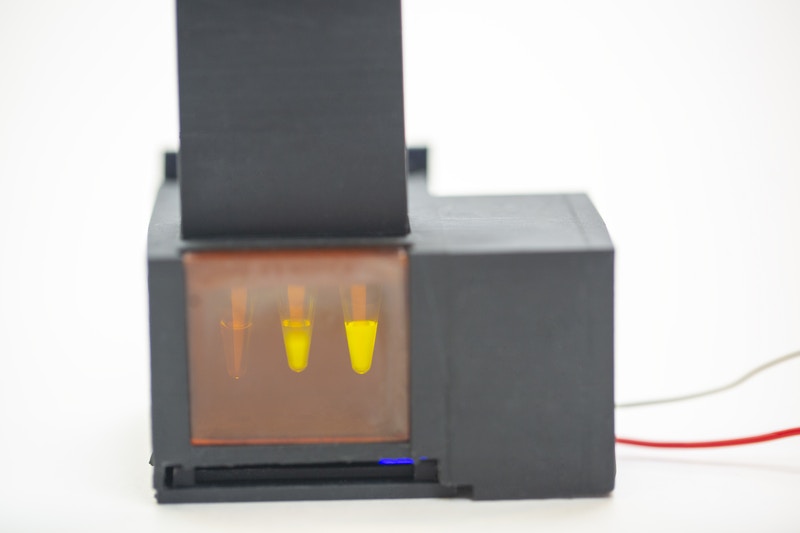
The saliva sample prep and the SHERLOCK reaction are integrated using a simple battery-powered device that has a heated sample prep chamber and an unheated reaction chamber.
A user spits into the sample preparation chamber, turns on the heat and has to wait three to six minutes for the saliva to be “wicked” into the filter. The user then removes the filter, transfers it to the reaction chamber column and then pushes a plunger that sends the filter into the chamber while puncturing a water reservoir to activate the SHERLOCK reaction. After about 55 minutes, the user looks into the reaction chamber through a tinted transilluminator window to see whether there is a fluorescent signal, which indicates a positive result. The accompanying smartphone app can also be used to analyze images taken from a smartphone camera to determine result positivity or negativity.
SHERLOCK Design: CRISPR COVID-19 Test
The research group developed the diagnostic using a CRISPR-based technology called “specific high sensitivity enzymatic reporter unlocking” (SHERLOCK), which was created in the lab of Dr. Jim Collins, Wyss Core Faculty member and senior paper author.
SHERLOCK is based on the CRISPR “molecular scissors” system involving a Cas enzyme that cuts DNA or RNA at specific locations, but with an additional snippet: upon recognizing its target sequence, the specially designed scissors also cut single-stranded engineered fluorescent DNA probes in the surrounding area. When these probes are cut along with the target sequence, a fluorescent signal is produced, indicating that the target has been successfully cut.

This “SHERLOCK reaction” is designed to cut SARS-CoV-2 RNA at a specific region in the nucleoprotein gene that is conserved across multiple variants of the virus, and which codes for proteins involved in formation of the viral nucleocapsid. The researchers also created additional SHERLOCK assays designed to target a panel of viral mutations in spike protein sequences that represent three SARS-CoV-2 genetic variants: Alpha, Beta and Gamma.
While several CRISPR-based tests have been developed for the detection of SARS-CoV-2, the workflows typically involve multiple liquid-handling steps and a plethora of lab equipment such as centrifuges and heating blocks, along with the technical skill to perform the procedures. This is labor-, time- and cost-intensive.
In contrast, miSHERLOCK has a modular design with components that can be assembled using a 3D printer and other common, inexpensive parts (around $15). The hardware, which can also be made with a 3D printer, can be re-used, bringing the cost of an individual assay to just $6. Traditional lab-based PCR tests can run anywhere between $20 and upwards of $500.
“One of the great things about miSHERLOCK is that it’s entirely modular. The device itself is separate from the assays, so you can plug in different assays for the specific sequence of RNA or DNA you’re trying to detect,” said co-first author Devora Najjar, a research assistant at the MIT Media Lab and in the Collins Lab, in a news release from the Wyss Institute. “The device costs about $15, but mass production would bring the housing costs down to about $3. Assays for new targets can be created in about two weeks, enabling the rapid development of tests for new variants of SARS-CoV-2 as well as for other infectious diseases.”
The researchers evaluated miSHERLOCK using clinical saliva samples from 27 COVID-19 patients and 21 healthy patients, finding that the diagnostic correctly identified COVID-19-positive patients 96 percent of the time and individuals without the disease 95 percent of the time. The sample size in this evaluation was small and thus further studies involving larger sample cohorts are warranted.
They also looked to see how the CRISPR COVID-19 test fared in detecting the Alpha, Beta, and Gamma SARS-CoV-2 variants by spiking healthy human saliva with full-length synthetic viral RNA containing mutations representing each variant, finding that the device could differentiate between the three variants and was effective across a range of viral RNA concentrations.
“When the miSHERLOCK project started, there was almost no SARS-CoV-2 variant monitoring happening. We knew that variant tracking was going to be incredibly important when evaluating the long-term effects of COVID-19 on local and global communities, so we pushed ourselves to create a truly decentralized, flexible, user-friendly diagnostic platform,” said Dr. Collins.









Join or login to leave a comment
JOIN LOGIN